|
1/26/2024 0 Comments 4peaks tutorial
Once your reverse complement sequence is available in the “Tool Output” window of EMBOSS Seqret, copy the whole sequence into the new text editor document (Ctrl + C and Ctrl + V). Open an empty text editor document on your computer.ħ. To receive the reverse complement of the sequence, click on “More options” and select “Yes” as “Reverse” option.Ħ. In “Step 2” select “FASTA format” as input and output format. In “Step 1” ensure “DNA” is selected as input data. Alternatively, use the upload function to upload the reverse. 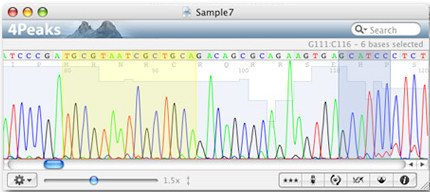
Paste all the information into the EMBOSS Seqret input box below (Ctrl + V). Copy the whole sequence including the “>” sign and descriptive header (keyboard shortcut Ctrl + C).ģ. seq file contains the sequence information in FASTA format.Ģ. Open the .seq file of your reverse sequence using a text editor such as NotePad on Windows or TextEdit on Mac. converting the 3′-5′ sequence into a 5′-3′ orientation). This can be achieved by converting the reverse sequence into its reverse complement (i.e. To be able to align the forward and reverse sequence reads, the two sequences have to have the same orientation. In case you would like to be guided through the instructions in more detail, please visit the Bioinformatics Tutorial page. To obtain your contig, follow the instructions in the tabs below. 
editing and assembling the consensus sequence aligning the forward and reverse sequence readsģ. converting the reverse sequence into its reverse complementĢ. Assembling the contig of the DNA barcode will involve the following steps:ġ. Before you can search the barcode against the entries in the European Nucleotide Archive (ENA), the sequences of the forward and reverse reads have to be assembled into a single consensus sequence called a contig. You have now received a forward and reverse sequence of your plant DNA barcode from the sequencing service. The analysis includes preparing the sequencing sample for database search and the database search itself, and will lead to identification of the plant species in question on a molecular basis. The following instructions guide through the analysis of the sequencing data of the DNA barcoding marker gene(s) which were obtained by DNA extraction and amplification in the wet-lab aspects of the DNA barcoding workflow.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
